The concept of transgenerational inheritance and genetic memory is one of the more intriguing aspects in epigenetic research. It’s also sparked more than a little debate and controversy about how humans and other organisms pass on genetic information. Check out this excerpt from Tom Brock’s article, that takes a deeper dive into the matter:
Genetic Memory
Is anything forever? Can anything truly be immortal on a planet that might end in 2012 (Mayan date) or be uninhabitable in 2.3 billion years?¹ In a similar way, how stable is our genome? Australopithecines, like Lucy and Ardi, walked the earth just 3 to 4 million years ago, and neanderthals were foraging in Eurasia 100,000 years ago. Certainly, something better than Homo sapiens is likely to emerge in the (relatively) near future. This suggests that DNA, itself, is constantly changing, adapting, taking a shape that bears witness to the need to adjust to evolving conditions. Offspring certainly resemble their parents and even their grandparents, attesting to the persistence of genotype for a few generations. However, the regular shuffling of the DNA deck during sexual reproduction, combined with an inherent error rate in DNA duplication during mitosis and a varying rate of mutation, indicates that our DNA is but a faint memory of our ancestors and their experiences.
Download this full article, Sex, Immortatlity and Genetic Memory by Tom Brock, Ph.D compliments of our friends at Cayman Chemical.
In a similar way, epigenetic marks are limited in their persistence. The most stable marks, DNA methyl groups, are completely erased during meiosis, only to be reestablished shortly after. While there are DNA methyltransferases that are dedicated to maintaining these marks on cytosines, newly synthesized DNA strands are produced with unmethylated cytosine bases. This means that marks must be actively re-established following DNA replication. Conceptually, there must be mechanisms in place to faithfully reproduce methyl marks on new DNA, as well as checking, correcting, and repairing processes akin to those that work on DNA. New experiences, stresses, and signals trigger the placement of new marks, and there must be ways to decide whether to preserve these ‘memories’ by reproducing the marks following DNA replication. Thus, DNA methylation appears to be a way to adjust gene expression in a relatively stable way, long-term. Importantly, these adjustments can still be reversed, either by removing or simply not renewing marks.
If DNA methylation provides long-term memory to gene expression, then histone modification might represent short-term memory. The variety of enzymes that attach or remove marks suggests that acetylation and methylation may be used in many different systems to produce a relatively temporary change in gene expression in specific ways. For example, the histone deacetylase HDAC2 participates in a cluster of proteins that acts as a functional model to maintain embryonic stem cell pluripotency.2 HDAC2, combined with Nanog, Oct4, and other interacting proteins (Figure 1), work in concert to mediate transcriptional repression that defines embryonic stem cells.
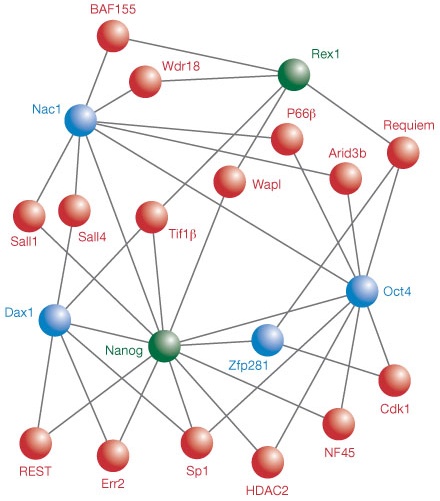 Stem cell marker proteins Nanog and Rex1 (green) were used to pull down interacting proteins, including core (blue) and peripheral (red) targets. (Adapted from Reference 2)
Stem cell marker proteins Nanog and Rex1 (green) were used to pull down interacting proteins, including core (blue) and peripheral (red) targets. (Adapted from Reference 2)
The beauty of histone modification is its flexibility. It can help a cell ‘remember’ that it is a stem cell for many cell cycles. Alternatively, specific signals can lead to a wholesale switch in how histones are marked, initiating cell differentiation.3,4 Changed histone marks serve to preserve the memory of the new cues while at the same time altering DNA utilization in the production of an intermediate blast-type cell. When the cell finally becomes, say, a FoxP3+ T cell, histone marks will not only play a central role in defining gene expression, they will be telltale clues to the cell’s history.
You can find the entire article, Sex, Immortatlity and Genetic Memory by Tom Brock Ph.D. on the Cayman Chemical Website.
References
- Li, K.F., Pahlevan, K., Kirschvink, J.L., et al. Proc. Natl. Acad. Sci. USA 106, 9576-9579 (2009).
- Wang, J., Rao, S., Chu, J., et al. Nature 444, 364-368 (2006).
- Spivakov, M. and Fisher, A.G. Nat. Rev. Genet 8, 263-271 (2007).
- Vincent, A. and Van Seuningen, I. Differentiation 78, 99-107 (2009).
