Small Non-coding RNA Enrichment Can Be RISCy
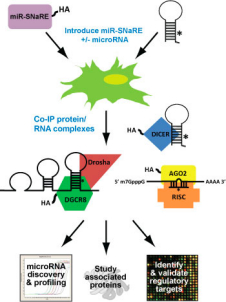
Things can get complicated when you are looking for low copy miRNAs, validating miRNA targets or trying to figure out all of the players in RISC complexes. SBI’s miR-SNaREs™, enable you to lasso those answers easily. The system uses Lentivectors to overexpress different flavors of epitope tagged microRNA processing factors that make pulling down DGCR8 (Pasha), DICER, AGO2 (Argonaute) and the RNAs (mRNA and ncRNA) they’re hanging out with a snap.
Constructs are available with co-expression of either RFP for fluorescent cell sorting or Puro for selection of stable cell lines.
What Next?
Hey what you do with your IPs is your biz, but you might want to consider:
- Cloning and sequencing the RNAs for ncRNA discovery
- Profiling miRNA that are being utilized in the RISC complex
- Search for other proteins that may be in on the action
- Validate some of the thousands of computational miRNA target predictions out there
Need More Info
Hey we don’t blame you. Good thing the folks at SBI have all the details, fancy graphics, and literature on their site. Check it out for yourself! Tell them we sent you.