DNA methylation has been a cornerstone of epigenetics research from the start, and the emergence of technologies that make investigation of the modification more accessible has only made the DNA methylation field even hotter. Several tools and techniques are available to assess DNA methylation levels, but two in particular, reduced representation bisulfite sequencing (RRBS) and the Infinium HumanMethylation450 BeadChip Array (450k array), have become increasingly popular due to their ability to deliver site specific detail at a genome-wide scale.
The Heyday of the 450k Array
Historically, the 450k array has been the more popular platform for DNA methylation analysis. Familiar array based protocols, simpler data analysis, and relatively lower costs as compared to sequencing, established the 450k arrays as a highly useful tool for analyzing differentially methylated regions (DMRs), especially when studying known CpG sites in human samples.
However, the 450k array does have some inherent problems and limitations. The arrays are spotted with pre-designed probes that correspond to human CpG locations. That presents a problem for incorporation of genetic information like single nucleotide polymorphism (SNP) detection, discovery of new methylation loci, or working with other species. In addition, the array probes are susceptible to batch effects and can potentially cross-hybridize with non-targeted DNA, confounding results and requiring confirmation of data using a second method.
RRBS Rising
RRBS is a powerful method for genome-wide DNA methylation analysis that focuses bisulfite sequencing to the CpG-rich regions where DNA methylation is most frequent, which saves both time and money. This approach utilizes restriction enzymes that digest CpG motifs, and the regions containing CpGs are recovered by gel purification. Genomic regions that don’t have many CpG sites remain large in size after the digestion, and are therefore not present in the gel-purified samples. The CpG-enriched DNA samples are bisulfite converted and further prepped for next-generation sequencing (NGS). Despite being developed as a way to minimize NGS costs, RRBS is still often perceived as expensive and difficult to perform, but falling sequencing prices, enhanced services, and intuitive bioinformatics have fueled a rise in RRBS accessibility.
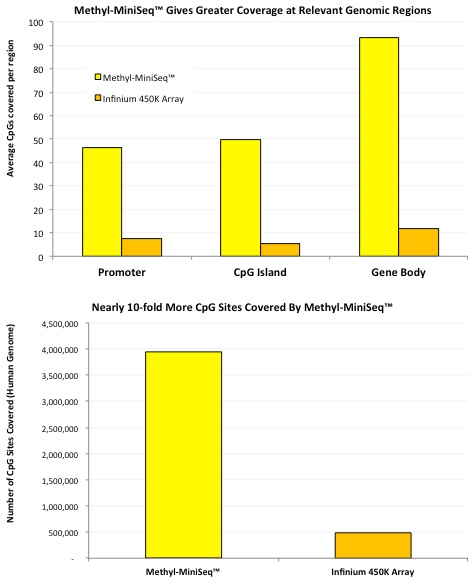
The methylation moguls at Zymo Research did some head-to-head comparisons (see figures below) while developing their RRBS-based Methyl-MiniSeq™ service and found that RRBS brings some pretty hefty benefits to the table, including:
- Compatibility with any organism that has a reference genome.
- Many more sites are covered, and at a greater depth; including more sites covered in important genomic regions like promoters, CpG islands, and gene bodies.
- The NGS component can simultaneously detect DNA methylation and SNPs, enabling both genetic and epigenetic analysis.
- On a cost-per-site basis, RRBS is likely less expensive than 450k arrays.
These days, with the advent of services like Methyl-MiniSeq™ from Zymo Research, which includes everything from library prep to complete, and customizable bioinformatics, it’s even possible to have RRBS analysis done without any heavy lifting on your part.
Now that you’ve seen the pros and cons for both the RRBS and 450k array methods, it is time to decide which approach is best for you and your needs.