While most studies on the N6-methyladenosine (m6A) RNA modification have been on the straight and narrow, the research trajectory taken by the labs of Cosmas C. Giallourakis and Alan C. Mullen will have you running around in circles! More specifically, Chan Zhou and Benoit Molinie (along with their tight circle of friends) sought to discover whether what we know about linear m6A-modified mRNA holds true for the circular form of RNA known as circRNA.
Created by covalent linkage of the 3′ and 5′ ends of spliced RNA transcripts, circRNAs can interact with transcriptional machinery and microRNAs, and may even find use as new disease biomarkers. However, we still lack any description of the m6A modification (or lack thereof) and its consequences for these curvy RNAs.
The team first depleted ribosomal RNA, degraded linear RNA, and then used m6A RNA precipitation to discover the abundant m6A modification of “leftover” circRNAs from human embryonic stem cells (hESCs). Next, they applied a newly developed computational pipeline called “AutoCirc” to establish a transcriptome-wide map of m6A-modified circRNAs.
So just how did this study “shape” up?
- siRNA knockdown indicated that the m6A modification of circRNAs requires the actions of the METTL3/METTL14 synergistic complex, just like m6A modification of mRNAs
- When compared to other pipelines (CIRCexplorer and MapSplice), AutoCirc detected similar expression patterns in a quicker time (10x faster and 250x faster, respectively), while consuming fewer computing resources
- Comparisons indicated higher m6A levels for circRNAs composed of long single exons (more abundant), compared to multi-exon circRNAs (less abundant)
- Interestingly, the m6A-modification of circRNAs did not correlate with the m6A status of their corresponding mRNAs, as demonstrated by the 59% of m6A-modifieid circRNAs produced from exons that lacked correlating m6A-modified mRNA
- Comparison of m6A-modified circRNA profiles from hESCs with those found for HeLa cells underlined cell-specific patterning and abundance of m6A-modified circRNAs
- The m6A modification typically promotes mRNA degradation; however, m6A levels on circRNAs from both HeLa cells and hESCs positively correlated with circRNA expression levels
- Analysis of proteins known to interact with the m6A modification on mRNAs indicated that both YTHDF1 (promotes translation) and YTHDF2 (promotes decay) also interacted with m6A-modified circRNAs
- While m6A-modified mRNAs exhibit a shorter half-life than non-modified mRNAs, m6A-modified mRNAs from the same exons as m6A-modified circRNAs have an even shorter half-life that is regulated by YTHDF2
Now that this exciting study has identified and described m6A-modified circRNAs and uncovered some novel findings, we now need to further probe the consequences of this modification and how it can influence mRNA, or other RNA species, and regulate transcription.
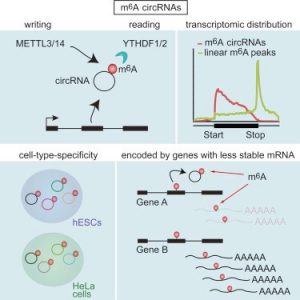
Graphical Abstract for Zhou et al
While we wait for the results of such studies, and for the (hopefully) soon to be discovered m6A-modification of square-, triangular-, and Christmas-tree-shaped-RNAs (!), satisfy yourself with a deeper dig into this well-rounded study at Cell Reports, August 2017.