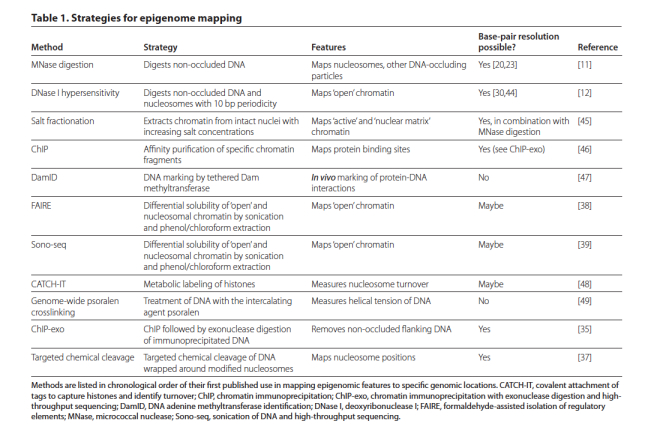
The days of dealing with regional estimates of epigenetic marks like histone modifications and DNA methylation are just about behind us. That’s right, the maps that lie ahead will be less Thomas Guide and more GPS. Drs. Gabriel Zentner and Steve Henikoff recently took some time off from being all-star cancer researchers and pulled together a nice review of some ways to resolve epigenetics signatures down to the base. Some of the methods were new to us, while others are re-mixes on old classics. Either way, the review is a full a good summary of some well-tested methods.
The Fred Hutch duo spent a little extra time on three methods that are based on familiar techniques, but with a twist. And they all involve DNA digestion. Here’s some of the info:
- MNase-seq. Researchers mainly use micrococcal nuclease (MNase) to digest chromatin to study nucleosomes, but they are also now using it to study non-histone proteins. These days, researchers are combining MNase digestion with paired-end sequencing and new ways of making libraries. This is called “MNase-seq,” and it can map a wide range of fragment sizes, localize lots of chromatin-bound proteins, and is cost-effective.
- DNase-seq. DNase-seq is DNase I digestion combined with high-throughput DNA sequencing to get sites of digestion at single base-pair resolution. It’s kind of the “yin” to MNase-seq’s “yang”, providing inferred info on chromatin-binding proteins that lie between hypersensitive sites, whereas MNase-seq maps protected regions. Both methods require another technique, such as ChIP-seq, for identifications of bound elements, though.
- ChIP-exo. ChIP is one of the most popular epigenomic strategies out there. But to get the method down to base-pair resolution, researchers have combined it with lambda exonuclease treatment. Mapping the boundaries of exonuclease activity allows for higher res than even ChIP-seq provides.
With a few tweaks, other classical methods also could provide base-pair resolution of epigenomic features. Check out the review for more details at: Genome Biology, October 2012.