Genome editing refers to an emerging branch of biotechnology that is the realization of earlier genetic engineering technologies. Using these biotechnologies researchers are able to target specific DNA sequences and induce a double stranded break, taking advantage of recombination to create synthetic genetic content in a host genome.
One of the way researchers are uncovering the secrets of epigenome is by utilizing genome editing tools, such as CRISPR-Cas9, ZFNs and TALENs, to modify key epigenetic players. For instance, researchers have taken the CRISPR-Cas9 system and used it to deplete key writers (Dmnt1, Dnmt3a, Dmnt3b) and readers (MeCP2) of the epigenome via indels in the adult mouse brain (Swiech et al. 2014). The approach was optimized to allow for efficient singleplex or multiplex targeting in the brain and was able to hit post-mitotic neurons, while also creating a great visualization system for successful transformation.
In addition, Musa Mhlanga and team were able to elucidate how chromatin looping regulates multi-gene complexes by using TALENs to alter the sequence in contact points critical to inter- and intra-chromosomal relations (Fanucchi et al., 2014). Finally, genome editing has enabled loss of function studies in human cells; for example, CRISPR/Cas9 showed that DNMT1 loss is lethal in human but not mouse embryonic stem cells (Liao et al., 2015).
Epigenome Editing
One way to overcome the challenges of studying the epigenome systematically is the use of manipulated genome editing tools. This approach is known as epigenome editing and relies on altering gene expression without altering the underlying sequences and is thus truly epigenetic. Epigenome editing systems rely on the targeting capabilities of a genome editing system, but are modified to have the nuclease inactivated or removed so it is incapable of cleaving DNA, but can still reach its target.
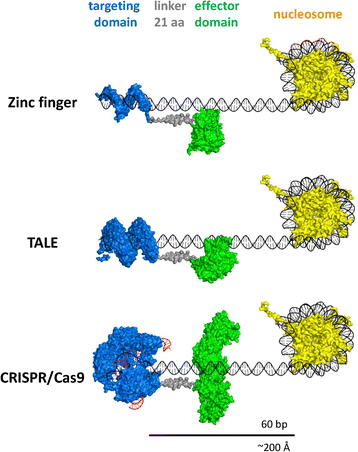
The novel functionality is given by a fusion of the altered genome editing system to an effector domain, which is a modular domain that gives the system the power of a transcription factor or epigenome modifier (Figure 1). Thus removing or inactivating the nuclease and adding an effector domain to provide new function confers the modular powers of a swiss army knife and allows for manipulation of the epigenome in several different ways.
Figure 1. Structural models of possible epigenetic editing devices. Figure reproduced from Jurkowski et al. 2015, Licenced under CC By 4.o by BioMed Central.
Artificial Transcription Factors (ATFs)
Artificial transcription factors have long been a dream of both the clinician and basic scientist. Essentially they are designer transcription factors that are capable of recruiting or keeping transcriptional machinery away from a specific sequence. Before being manipulated to edit the genome, Zinc Finger Proteins (ZFPs) and Transcription Activator-Like Effectors (TALEs) biotechnologies were first discovered and utilized as (artificial) transcription factors. Now CRISPR-Cas9 has also been modified to act in this manner. ATFs act to either activate or repress transcription much like a biological factor, but with the power of synthetics, allowing for the functions of activating or repressing transcription.
Activating Transcription
The ability to artificially activate transcription came about with VP16, which is a 16 amino acid [DALDDFDLDML] peptide from the Herpes Simplex Virus (Seipel et al., 1992). It was later assembled into VP64, which is four copies of VP16 connected by glycine-serine linkers (Beerli et al., 1998). The first example of CRISPR-Cas9 system being hijacked to activate transcription was by researchers in J Keith Joung’s lab who used a VP64 transactivation domain fused to a deactivated Cas9 (dCas9) (Maeder et al., 2013). A study comparing the ability of dCas9 and TALEs to drive transcription showed that each system has context dependent merits when it comes to effect and recommended a combined approach (Gao et al., 2014). It also showed that VP64 can recruit p300 and result in H3K27ac, which is an example of transcription driving transcription. A more recent approach from the lab of Feng Zhang used an engineered synergistic CRISPR/Cas9 activator complex for genome-wide transcriptional control (Konermann et al., 2015). The utility of CRISPR-Cas9 to activate transcription has been highlighted by it’s successful application in vivo by scientists headed by Rudolf Jaenisch to alter developing zygotes (Cheng et al., 2013). Recently, George Church’s group has developed a tripartite activator utilizing a fusion of VP64-p65-Rta that targets in multiplex and can stimulate neuronal differentiation of human iPSCs (Chavez et al., 2015). These systems recruit transcriptional machinery to the target site and activate transcription, all while not changing the underlying targeted DNA sequence.
Repressing Transcription
In its first form CRISPR-Cas9 repressed transcription through utilizing steric hindrance to simply block the target site from transcription machinery. This system was further developed to utilize transcriptional repression domains, allowing for stronger repression than steric hindrance alone (Gilbert et al., 2013).
Editing the Epigenome
Altering gene expression in an inheritable fashion without changing DNA sequence is the epitome of epigenome editing. By taking some of our favorite epigenome writers and erasers and using them as the effector module, new powers can be conferred onto the existing targeting systems.
Modifying Histones
Epigenome editing has been used to figure out the function of histone modifications through targeting histone modifiers to specific regions of the genome. By fusing a Zinc Finger Protein to a histone methyltransferase researchers led by Carl Pabo have been able to show that H3K9 methylation can initiate gene repression (Snowden et al., 2002). In addition, researchers in the lab of Bradley Bernstein used TALEs fused to the LSD1 histone demethylase to remove histone modifications only at enhancer sequences and thus characterize their function (Mendenhall et al., 2013). The lab of Feng Zhang has created an optogenetic epiTALE system in which an inducible element both activates and represses transcription via effector domains, many of which modify histones (Konermann et al., 2014). In neurons, the levels of H3K9me1, H4K20me3, H3K27me3, H3K9ac, and H4K8ac were altered by epiTALEs made from a number of your favorite epigenetic enzymes. CRISPR has also joined the party: in one study, dCas9 was utilized with a p300 effector domain, creating a precision histone acetyltransferase that can overcome the challenges of activating enhancers (Hilton et al., 2015).
Demethylating Cytosine
Thymidine DNA glycosylase (TDG) was first to the scene and was used to remove repressive DNA methylation and induce gene expression (Gregory et al., 2013). The famous TET family has also been utilized in epigenome editing. This was pioneered by researchers in J Keith Joung’s lab who fused TALEs to the TET1 hydroxylase catalytic domain, allowing for targeted DNA demethylation in human cell lines (Maeder et al., 2013). In the clinical environment, this system has potential for targeting epigenetically silenced tumor suppressors. Arising to the challenge, a system using a TALE and TET-2 fusion was created in the lab of Marianne G. Rots and was shown to be able to target an epigenetically silenced cancer gene (ICAM-1) and induce its expression (Chen et al., 2014).
Non-coding RNA
CRISPR-Display takes advantage of an upgraded synthetic guide RNA (sgRNA) that contains 3 domains capable of precisely displaying ncRNAs when bound to dCas9 (Shechner et al., 2015). The domains are located at the 5’ end, the 3’ end, and the middle of the sgRNA. Also, the custom cargo can be anything from lncRNAs, protein-binding cassettes that allow for bridged activation, artificial aptamers, and pools of random sequences.
Even More FUNctions
The fun of a nuclease null system doesn’t end yet as these systems can also have functions that aid epigenetic research.
Locus Specific ChIP
dCas9 can be endowed with a handy tag that gives you the power to precisely pull down proteins and RNAs interacting with your sequence of interest (Fujita & Fujii, 2013). Check out this EpiGenie webinar for more.
Optogenetic Induction
Optogenetic activation is ready to go with dCas9 (Nihongaki et al, 2015 & Polstein & Gersbach, 2015). Briefly, optogenetics involves Light-sensitive cryptochrome 2 (CRY2) and binding partner CIB1. When blue light hits, the two proteins come together and form the bridge needed to give dCas9 some temporal powers.
Cell Imaging
By tagging GFP and other fluorescent molecules onto dCas9, the cell begins to light up precisely the way you wanted it to. This approach has been used in live human cells and mouse embryonic stem cells (Chen et al. 2013 & Anton et al. 2014)
Further Learning:
This comprehensive review introduces the use of effector domains in nuclease null genome editing systems, which bring about desired epigenetic or transcriptional effects.
Voigt P, Reinberg D. Epigenome editing. Nat Biotechnol. 2013 (12):1097–1099.
This brief review covers some of the first breakthroughs in modifying genome editing systems for epigenome editing and transcriptional control.
Introduction to Epigenetic Editing
An EpiGenie webinar with thought leader Dr. Marianne Rots on the current state and future of epigenetic editing.
Day JJ. New approaches to manipulating the epigenome. Dialogues Clin Neurosci 2014 Sep;16(3):345-357
This review covers epigenome editing and optogenetics as tools to understand the brain.
A clinically oriented review on synthetic epigenetics.
In this paper, dCas9 was fused to effector domains, both activating and repressing, in order to reprogram human embryonic stem cells and control their differentiation
A recent review of synthetic approaches to regulate chromatin and transcription in eukaryotes.
One of the first reviews of epigenome editing that covers seminal breakthroughs with a translational perspective.
A protocol for generating Transcription Activator-Like Effectors (TALEs)
A protocol for developing custom Zinc Finger Protein (ZFP) arrays that also covers their use as artificial transcription factors.
References:
Swiech L, Heidenreich M, Banerjee A, Habib N, Li Y, Trombetta J, et al. In vivo interrogation of gene function in the mammalian brain using CRISPR-Cas9. Nat Biotechnol 2015 33:102–6.
Fanucchi S, Shibayama Y, Burd S, Weinberg MS, Mhlanga MM. Chromosomal contact permits transcription between coregulated genes. Cell 2013 155:606–20.
Liao J, Karnik R, Gu H, Ziller MJ, Clement K, Tsankov AM, Akopian V, Gifford CA, Donaghey J, Galonska C, Pop R, Reyon D, Tsai SQ, Mallard W, Joung JK, Rinn JL, Gnirke A, Meissner A. Targeted disruption of DNMT1, DNMT3A and DNMT3B in human embryonic stem cells. Nat Genet. 2015 47:469–78.
Seipel K, Georgiev O, Schaffner W. Different activation domains stimu- late transcription from remote (‘enhancer’) and proximal (‘promoter’) positions. EMBO J. 1992 11:4961–8.
Beerli RR, Segal DJ, Dreier B, Barbas CF 3rd. Toward controlling gene expression at will: specific regulation of the erbB-2/HER-2 promoter by using polydactyl zinc finger proteins constructed from modular building blocks. Proc Natl Acad Sci USA. 1998 95:14628–33.
Gao X, Tsang JC, Gaba F, Wu D, Lu L, Liu P. Comparison of TALE designer transcription factors and the CRISPR/dCas9 in regulation of gene expression by targeting enhancers. Nucleic Acids Res. 2014 42:e155.
Maeder ML, Linder SJ, Cascio VM, Fu Y, Ho QH, Joung JK. CRISPR RNA-guided activation of endogenous human genes. Nat Methods 2013 10:977–9.
Konermann S, Brigham MD, Trevino AE, Joung J, Abudayyeh OO, Barcena C, et al. Genome-scale transcriptional activation by an engineered CRISPR-Cas9 complex. Nature 2015 517:583–8.
Cheng AW, Wang H, Yang H, Shi L, Katz Y, Theunissen TW, et al. Multiplexed activation of endogenous genes by CRISPR-on, an RNA-guided transcriptional activator system. Cell Res 2013 23:1163–71.
Chavez A, Scheiman J, Vora S, Pruitt BW, Tuttle M, P R Iyer E, et al. Highly efficient Cas9-mediated transcriptional programming. Nat Methods 2015. Advanced online publication .
Gilbert LA, Larson MH, Morsut L, Liu Z, Brar GA, Torres SE, et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 2013 154:442–51.
Snowden AW, Gregory PD, Case CC, Pabo CO. Gene-specific targeting of H3K9 methylation is sufficient for initiating repression in vivo. Curr Biol 2002 12:2159–2166.
Mendenhall EM, Williamson KE, Reyon D, Zou JY, Ram O, Joung JK, et al. Locus-specific editing of histone modifications at endogenous enhancers. Nat Biotechnol 2013 31:1133–6.
Konermann S, Brigham MD, Trevino AE, Hsu PD, Heidenreich M, Cong L, et al. Optical control of mammalian endogenous transcription and epigenetic states. Nature 2014 500:472-476.
Hilton IB, D’Ippolito AM, Vockley, CM, Thakore PI, Crawford GE, Reddy TE, & Gersbach CA. Epigenome editing by a CRISPR-Cas9-based acetyltransferase activates genes from promoters and enhancers. Nature Biotechnol 2015 33:510-7.
Maeder ML, Angstman JF, Richardson ME, Linder SJ, Cascio VM, Tsai SQ, et al. Targeted DNA demethylation and activation of endogenous genes using programmable TALE-TET1 fusion proteins. Nat Biotechnol 2013 31:1137–42.
Chen H, Kazemier HG, de Groote ML, Ruiters MH, Xu GL, Rots MG. Induced DNA demethylation by targeting Ten-Eleven Translocation 2 to the human ICAM-1 promoter. Nucleic Acids Res 2014 42:1563–74.
Shechner, DM., Hacisuleyman, E, Younger, ST, & Rinn, JL. Multiplexable, locus-specific targeting of long RNAs with CRISPR-Display. Nat Methods. 2015. 12:664-7
Fujita T, Fujii H. Efficient isolation of specific genomic regions and identification of associated proteins by engineered DNA-binding molecule-mediated chromatin immunoprecipitation (enChIP) using CRISPR. Biochem Biophys Res Commun 2013 439:132–6.
Nihongaki Y, Yamamoto S, Kawano F, Suzuki H, & Sato M. CRISPR-Cas9-based photoactivatable transcription system. Chem Biol 2015 22:169-174.
Polstein LR, & Gersbach CA. A light-inducible CRISPR-Cas9 system for control of endogenous gene activation. Nat chem Biol. 2015 11:198-200.
Chen B, Gilbert LA, Cimini BA, Schnitzbauer J, Zhang W, Li GW, Park J, Blackburn EH, Weissman JS, Qi LS, Huang B. Dynamic imaging of genomic loci in living human cells by an optimized CRISPR/Cas system. Cell 2013 155:1479–91.
Anton T, Bultmann S, Leonhardt H, Markaki Y. Visualization of specific DNA sequences in living mouse embryonic stem cells with a programmable fluorescent CRISPR/Cas system. Nucleus 2014 5:163–72.