Imagine a world where you could create chromatin loops where and when you want. You’d feel like you’re on cloud nine. Well, get your head out of the clouds, because you’ve got to thank the lab of Kevin Wang at Stanford for targeted and reversible chromatin loop reorganization using deactivated Cas9 (CLOuD9).
The CLOuD9 system makes use of two dCas9 constructs, one from S. aureus and the other from S. pyogenes, where one half of a dimerization construct is tethered to the C-terminus of a dCas9. Then, abscisic acid (ABA) is added to dimerize the proteins and create a chromatin loop. The system is reversible, which can be accomplished by removing ABA via ligand washout, and can also be targeted to any locus with the right sgRNAs.
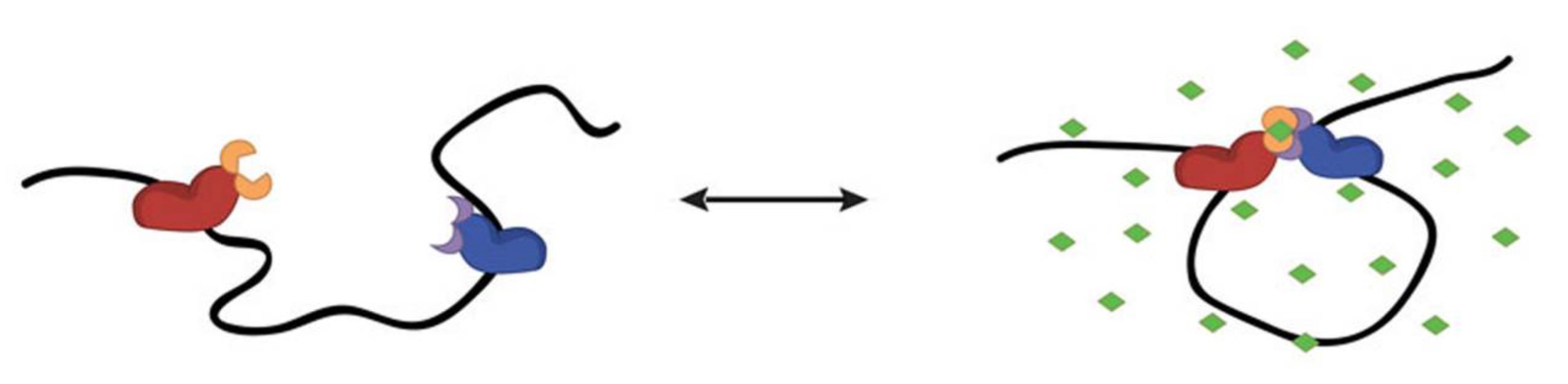
Schematic of artificial looping from Morgan et al., where ABA (green) brings complimentary CLOuD9 constructs (red and blue) together and removal of ABA restores the original chromatin conformation.
The team primarily targeted the locus control region (LCR) of the human β-globin gene, which regulates the expression of distant globin genes through a long-range chromatin loop. They turned to the K562 human erythroleukemia cell line to see if they could re-establish β-globin gene expression by bringing the LCR closer to the β-globin gene. Since the locus is in a large region of euchromatin, they also wanted to see what happened when they applied their construct heterochromatin, so they chose HEK 293T cells, where the globin genes aren’t expressed.
Here’s what went down:
- At first the two cell-types seemed similar, chromosome conformation capture (3C) revealed that in both cell types:
- just 24 hours after the addition of ABA there are increased contacts between the two loci
- the contacts are reversible and this happens as quickly as 24 hours after ligand washout
- Despite both cell types showing increased contacts, qPCR uncovered the difference between the two cell types, where only K562 cells showed increased:
- gene expression via RT-qPCR
- activating H3K4me3 and RNA Pol II occupancy by ChIP-qPCR
- After a 10-day ABA binge, K562 cells become stable and no longer show a reversal of the chromatin contacts or gene expression after ligand washout.
- Mass spectrometry analysis following ChIP (MS/ChIP) uncovered that RNA helicases (DDX5 and DDX17) are enriched for and remain after ligand washout, while CTCF and cohesin are surprisingly absent from the party
- While shRNA knockdown of both RNA helicases resulted in cell death, the individual knockdowns only interfered with the stability of the long-term chromatin contacts and gene expression following ligand washout
Overall, this study provides a tool that allows for you to selectively alter gene expression by artificial chromosome loops in either a reversible and stable fashion. On top of all that, it also offers new insights into the players behind chromatin loop function and stability.
Go learn how to put your favorite sequence on CLOuD9 over at Nature Communications, July 2017