In Greek mythology, the fates forge the strings of life that control our lives and choose the moment of its end. We mere mortals yearn for such powers and we are now one step closer to the light thanks to an optogenetic epigenome editing system that lets you write and erase DNA methylation to control stem cell fate. This legendary achievement stems from a collaboration between the labs of Joseph Irudayaraj at Purdue University and Feng C. Zhou at Indiana University.
Previous studies accomplished optogenetic epigenetic editing with a Transcription Activator-Like Element (TALE) via histone effectors with DNA methylation recently joining the light show. This strategy involves fusing a TALE to CRY2 and the effector, a catalytic domain of choice, to CIB1. Blue light then dimerizes the optogenetic protein pairs (CRY2 and CIB1) and unleashes the precision epigenetic editing system. In terms of effector domains, the DNMT3A catalytic domain writes DNA methylation, while the TET1 catalytic domain helps to erase DNA methylation.
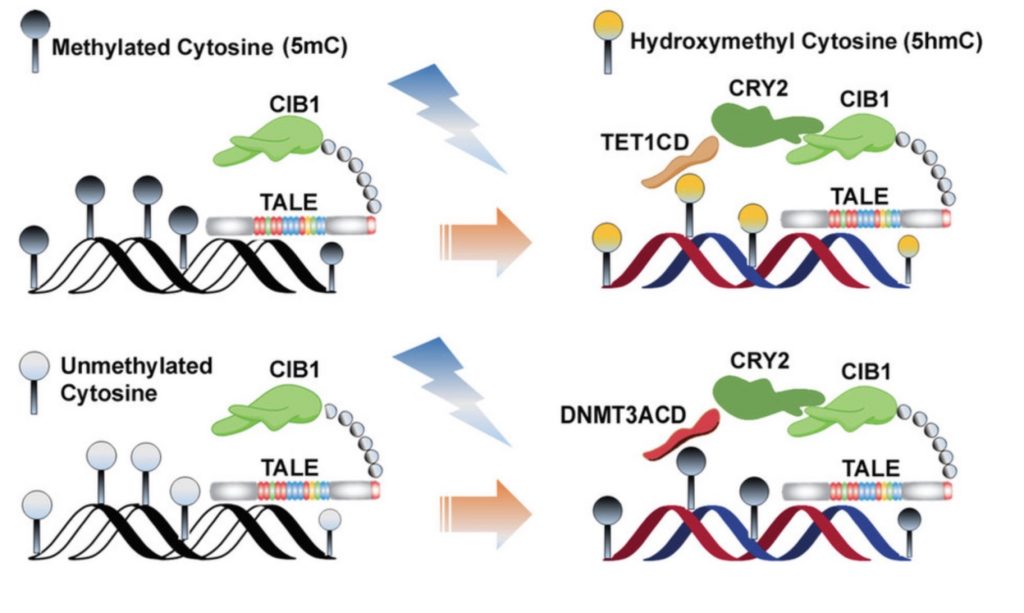 Optogenetic editing of DNA methylation (CC BY 4.0)
Optogenetic editing of DNA methylation (CC BY 4.0)
In this case, the team designed a TALE to target their effector domains to the promoter of Ascl1, which determines whether neural stem cells (NSCs) develop into neurons or glia. Given their divergent phenotypic fates, the team manipulated rat dorsal root ganglion (DRG) and striatal (STR) NSCs.
To read destiny, Lo et al. used bisulfite amplicon sequencing to investigate the methylation of Ascl1 and 300 bp of its promoter sequence. They chose to target a 30 bp region within the promoter since it contained 4 CpGs with multiple transcription factor binding sites and confirmed the methylation differences there by pyrosequencing. They also used RT-qPCR to relatively quantify Ascl1 expression.
Here’s what went down:
- They confirmed that Ascl1 is hypomethylated in DRG NSCs, which preferentially generate neurons, and hypermethylated in STR NSCs, which preferentially generate glia.
- Demonstrating the regulatory nature of methylation, they found higher Ascl1 expression levels in the hypomethylated DRG NSCs.
- They also found that as STR NSCs differentiate methylation levels decrease and expression increases.
- By inducing higher levels of DNA methylation with the DNMT3A catalytic domain in DRG NSCs, the team decreased gene expression.
- Promoting DNA hydroxymethylation with the TET1 catalytic domain in STR NSCs led to the erasure of DNA methylation and increased gene expression.
Moving from function to phenotype, they examined the neuronal marker NeuN with immunocytochemistry. This revealed that STR NSCs transfected with the TET1 construct showed higher NeuN protein expression, and thus had their fates shifted towards neurons rather than the typical glial destinies.
Overall, this thought-provoking study demonstrates how epigenetic editing can establish the functional roles of regulatory elements, while also controlling phenotype destiny via an epigenetic switch that lets you play with the very fabric of NSC fate.
Go learn how to shine some light on the fate of your epigenetic editing experiments in Scientific Reports, February 2017
